Your new post is loading...
 Your new post is loading...
Authors: Sahana Basu, Monika, Surbhi Kumari and Gautam Kumar.
Plant Physiology and Biochemistry (2024)
Highlights: • IAA induces lateral root formation in rice under submergence • ROS and RNS cause lysigenous aerenchyma formation in root cortex of rice • Submergence decreases ABA level, causing GA-mediated escape response in rice shoot • Enhanced antioxidant levels dampen nitro-oxidative damage during de-submergence
Abstract: "Constant change in global climate has become the most important limiting factor to crop productivity. Asymmetrical precipitations are causing recurrent flood events around the world. Submergence is one of the most detrimental abiotic stresses for sustainable rice production in the rainfed ecosystems of Southeast Asia. Therefore, the development of submergence-tolerant rice is an essential requirement to encounter food security. Submergence tolerance in rice is governed by the major quantitative trait locus (QTL) designated as Submergence1 (Sub1) near the centromere of chromosome 9. The introduction of the Sub1 in high-yielding rice varieties producing near-isogenic lines (NILs) has shown extreme submergence tolerance. The present study aimed to understand the responses of rice genotype IR64 and its Sub1 NIL IR64 Sub1 following one week of complete submergence treatment. Submergence imposed severe nitro-oxidative stress in both the rice genotypes, consequently disrupting the cellular redox homeostasis. In this study, IR64 exhibited higher NADPH oxidase activity accompanied by increased reactive oxygen species, reactive nitrogen species, and malondialdehyde buildups and cell death under submergence. Higher accumulations of 1-Aminocyclopropane-1-carboxylic acid, gibberellic acid, and Indole-3-acetic acid were also observed in IR64 which accelerated the plant growth and root cortical aerenchyma development following submergence. In contrast, IR64 Sub1 had enhanced submergence tolerance associated with an improved antioxidant defense system with sustainable morpho-physiological activities and restricted root aerenchyma formation. The comprehensive analyses of the responses of rice genotypes with contrasting submergence tolerance may demonstrate the intricacies of rice under complete submergence and may potentially contribute to improving stress resilience by advancing our understanding of the mechanisms of submergence tolerance in rice."
Authors: Tao Zhang, Xinyu Wang, Yanchao Yuan, Shoujie Zhu, Chunying Liu, Yuxi Zhang, Shupeng Gai.
Horticulture Research (2024)
Abstract: "Bud endodormancy in perennial plants is a sophisticated system that adapts to seasonal climatic changes. Growth-promoting signals such as low temperature and gibberellins (GAs) are crucial for facilitating budbreak following endodormancy release (EDR). However, the regulatory mechanisms underlying GA-mediated budbreak in tree peony (Paeonia suffruticosa) remain unclear. In tree peony, the expression of PsmiR159b among three differentially expressed miR159 members was inhibited with the prolonged chilling, and overexpression of PsMIR159b delayed budbreak, whereas silencing PsmiR159b promoted budbreak after dormancy. PsMYB65, a downstream transcription factor in the GA pathway, was induced by prolonged chilling and exogenous GA3 treatments. PsMYB65 was identified as a target of PsmiR159b, and promoted budbreak in tree peony. RNA-seq of PsMYB65-silenced buds revealed significant enrichment in the GO terms regulation of ‘cell cycle’ and ‘DNA replication’ among differentially expressed genes. Yeast one-hybrid and electrophoretic mobility shift assays demonstrated that PsMYB65 directly bound to the promoter of the type-D cyclin gene PsCYCD3;1. Dual-luciferase reporter assay indicated that PsMYB65 positively regulate PsCYCD3;1 expression, suggesting that miR159b-PsMYB65 module contributes to budbreak by influencing the cell cycle. Our findings revealed that the PsmiR159b-PsMYB65 module functioned in budbreak after dormancy by regulating cell proliferation, providing valuable insights into the endodormancy release regulation mechanism."
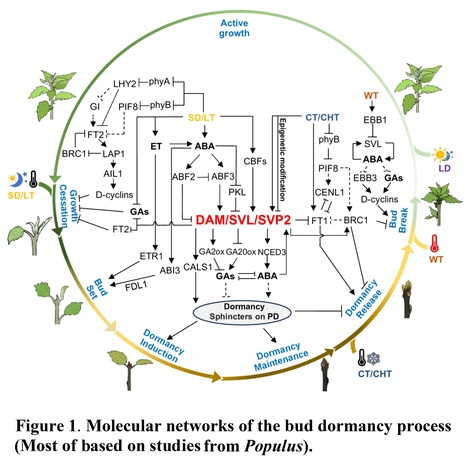
Authors: Jihua Ding, Kejing Wang, Shashank Pandey, Mariano Perales, Isabel Allona, Md Rezaul Islam Khan, Victor B. Busov, Rishikesh P. Bhalerao
Journal of Experimental Botany (2024)
Abstract: "Seasonal bud dormancy in perennial woody plants is a crucial and intricate process that is vital for the survival and development of plants. Over the past few decades, significant advancements have been made in understanding many features of bud dormancy, particularly in model species, where certain molecular mechanisms underlying this process have been elucidated. In this review, we provide an overview of recent molecular progress in understanding bud dormancy in trees, with a specific emphasis on the integration of common signaling and molecular mechanisms identified across different tree species. Additionally, we address some challenges that have emerged in the in-depth understanding of bud dormancy and offer insights for future studies."
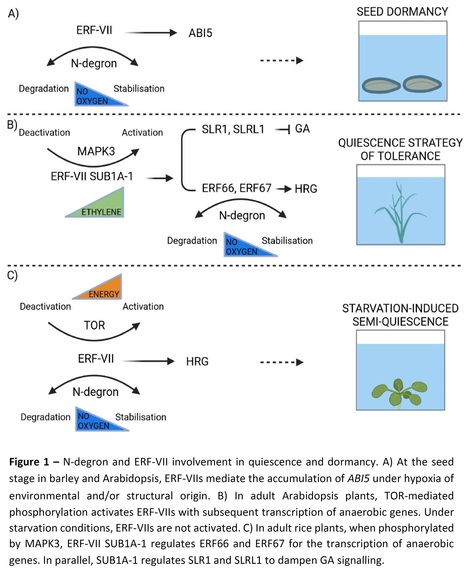
Authors: Chiara Pucciariello and Pierdomenico Perata.
Journal of Experimental Botany (2024)
Abstract: "Plant quiescence and seed dormancy can be triggered by reduced oxygen availability. Under water, oxygen depletion caused by flooding can culminate in a quiescent state, which is a plant strategy for energy preservation and survival. In adult plants, a quiescent state can be activated by sugar starvation, culminating in metabolic depression. In seeds, secondary dormancy can be activated by reduced oxygen availability, which creates an unfavourable state for germination. The physical dormancy of some seeds and buds includes barriers to external conditions, which indirectly results in hypoxia. The molecular processes that support seed dormancy and plant survival through quiescence under hypoxia include the N-degron pathway, which enables the modulation of ethylene responsive factors of group VII and downstream targets. This oxygen- and nitric oxide-dependent mechanism interacts with phytohormone-related pathways to control growth."
Authors: Vincent Castric, Rita A. Batista, Amélie Carré, Soraya Mousavi, Clément Mazoyer, Cécile Godé, Sophie Gallina, Chloé Ponitzki, Anthony Theron, Arnaud Bellec, William Marande, Sylvain Santoni, Roberto Mariotti, Andrea Rubini, Sylvain Legrand, Sylvain Billiard, Xavier Vekemans, Philippe Vernet and Pierre Saumitou-Laprade.
Current Biology (2024)
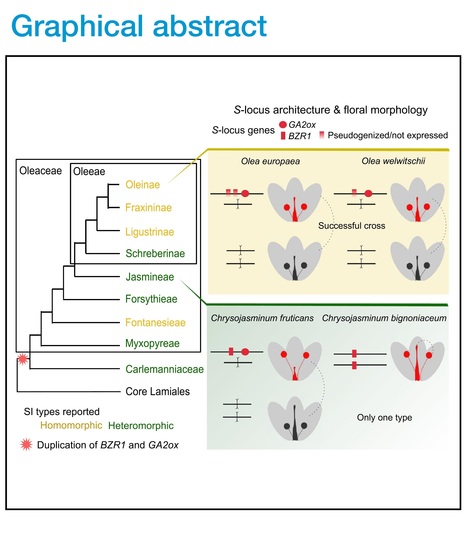
Editor's view: The self-incompatibility system of the Oleaceae is unusual, with the long-term maintenance of only two groups of reproductive compatibility. Castric et al. identify a hemizygous chromosomal fragment unique to one specificity, containing a single conserved gene with predicted function in catabolism of gibberellin.
Highlights: • The segregation of a genome fragment controls self-incompatibility (SI) in Oleaceae • This is the first report of hemizygous control of a homomorphic SI system • This fragment contains a single conserved gene, related to the gibberellin pathway • Manipulation of gibberellin levels switches self-incompatibility specificities
Abstract; "In flowering plants, outcrossing is commonly ensured by self-incompatibility (SI) systems. These can be homomorphic (typically with many different allelic specificities) or can accompany flower heteromorphism (mostly with just two specificities and corresponding floral types). The SI system of the Oleaceae family is unusual, with the long-term maintenance of only two specificities but often without flower morphology differences. To elucidate the genomic architecture and molecular basis of this SI system, we obtained chromosome-scale genome assemblies of Phillyrea angustifolia individuals and related them to a genetic map. The S-locus region proved to have a segregating 543-kb indel unique to one specificity, suggesting a hemizygous region, as observed in all distylous systems so far studied at the genomic level. Only one of the predicted genes in this indel region is found in the olive tree, Olea europaea, genome, also within a segregating indel. We describe complete association between the presence/absence of this gene and the SI types determined for individuals of seven distantly related Oleaceae species. This gene is predicted to be involved in catabolism of the gibberellic acid (GA) hormone, and experimental manipulation of GA levels in developing buds modified the male and female SI responses of the two specificities in different ways. Our results provide a unique example of a homomorphic SI system, where a single conserved gibberellin-related gene in a hemizygous indel underlies the long-term maintenance of two groups of reproductive compatibility."
Authors: Ting Li, Yongqin Wang, Annelore Natran, Yi Zhang, Hao Wang, Kangxi Du, Peng Qin, Hua Yuan, Weilan Chen, Bin Tu, Dirk Inzé and Marieke Dubois.
New Phytologist (2024)
Abstract: "Gibberellic acid (GA) plays a central role in many plant developmental processes and is crucial for crop improvement. DELLA proteins, the core suppressors in the GA signaling pathway, are degraded by GA via the 26S proteasomal pathway to release the GA response. However, little is known about the phosphorylation-mediated regulation of DELLA proteins. In this study, we combined GA response assays with protein–protein interaction analysis to infer the connection between Arabidopsis thaliana DELLAs and the C-TERMINAL DOMAIN PHOSPHATASE-LIKE 3 (CPL3), a phosphatase involved in the dephosphorylation of RNA polymerase II. We show that CPL3 directly interacts with DELLA proteins and promotes DELLA protein stability by inhibiting its degradation by the 26S proteasome. Consequently, CPL3 negatively modulates multiple GA-mediated processes of plant development, including hypocotyl elongation, flowering time, and anthocyanin accumulation. Taken together, our findings demonstrate that CPL3 serves as a novel regulator that could improve DELLA stability and thereby participate in GA signaling transduction."
Authors: Yangang Pei, Qihan Xue, Peng Shu, Weijie Xu, Xiaofei Du, Mengbo Wu, Kaidong Liu, Julien Pirrello, Mondher Bouzayen, Yiguo Hong and Mingchun Liu.
Developmental Cell (2024)
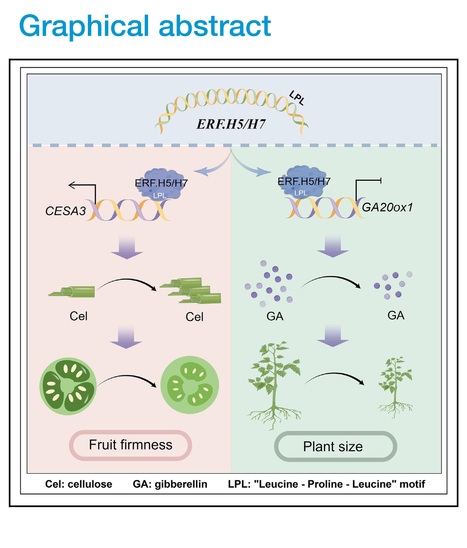
Editor's view: Pei et al. demonstrate the role of bifunctional transcription factors SlERF.H5 and SlERF.H7 in activating the cellulose biosynthesis gene SlCESA3 while repressing the gibberellin biosynthesis gene GA20ox1. The study highlights the distinct transcriptional regulatory functions of these ERFs in promoting cell wall formation and inhibiting plant growth.
Highlights • Cellulose is required for cell wall formation and firmness maintenance of fruits • SlERF.H5 and SlERF.H7 act as both transcriptional activators and repressors • The regulatory activity of SlERF.H5 and SlERF.H7 is mediated by a conserved motif
Abstract: "The plant cell wall is a dynamic structure that plays an essential role in development, but the mechanism regulating cell wall formation remains poorly understood. We demonstrate that two transcription factors, SlERF.H5 and SlERF.H7, control cell wall formation and tomato fruit firmness in an additive manner. Knockout of SlERF.H5, SlERF.H7, or both genes decreased cell wall thickness, firmness, and cellulose contents in fruits during early development, especially in double-knockout lines. Overexpressing either gene resulted in thicker cell walls and greater fruit firmness with elevated cellulose levels in fruits but severely dwarf plants with lower gibberellin contents. We further identified that SlERF.H5 and SlERF.H7 activate the cellulose biosynthesis gene SlCESA3 but repress the gibberellin biosynthesis gene GA20ox1. Moreover, we identified a conserved LPL motif in these ERFs responsible for their activities as transcriptional activators and repressors, providing insight into how bifunctional transcription factors modulate distinct developmental processes."
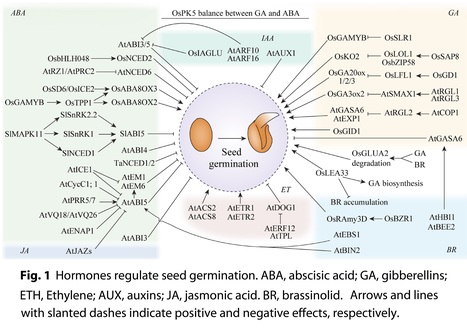
Authors: Jia Zhao, Yongqi He, Hongsheng Zhang and Zhoufei Wang.
Seed Biology (2024)
Abstract: "Seed germination is a key process in the life cycle of seed plants. The initiation of seed germination requires the activity of specific internal signaling molecules, such as hormones and reactive oxygen species (ROS), and is dependent on external environmental factors, such as water, temperature, and light. Seed germination is a complex trait that is regulated by multiple factors, including transcript, protein, and metabolite levels. This review highlights current knowledge relating to the regulatory roles of hormones, ROS, small RNAs, epigenetic modifications, post-translational modifications, and environmental cues on seed germination, mainly focusing on Arabidopsis and rice. The review on the molecular regulation of seed germination contributes to the improvement of crop seed quality using bio-breeding approaches."
Authors: Jennifer Andres, Lisa J. Schmunk, Federico Grau-Enguix, Justine Braguy, Sophia L. Samodelov, Tim Blomeier, Rocio Ochoa-Fernandez, Wilfried Weber, Salim Al-Babili, David Alabadí, Miguel A. Blázquez and Matias D. Zurbriggen.
The Plant Journal (2024)
Significance Statement: Using the degradation-based signaling mechanism of GAs, we have built transcriptional regulator (DELLA)-based, genetically encoded ratiometric biosensors as proxies for hormone quantification at high temporal resolution and sensitivity that allow dynamic, rapid and simple analysis in a plant cell system, i.e. Arabidopsis protoplasts.
Abstract: "Gibberellins (GAs) are major regulators of developmental and growth processes in plants. Using the degradation-based signaling mechanism of GAs, we have built transcriptional regulator (DELLA)-based, genetically encoded ratiometric biosensors as proxies for hormone quantification at high temporal resolution and sensitivity that allow dynamic, rapid and simple analysis in a plant cell system, i.e. Arabidopsis protoplasts. These ratiometric biosensors incorporate a DELLA protein as a degradation target fused to a firefly luciferase connected via a 2A peptide to a renilla luciferase as a co-expressed normalization element. We have implemented these biosensors for all five Arabidopsis DELLA proteins, GA-INSENSITIVE, GAI; REPRESSOR-of-ga1-3, RGA; RGA-like1, RGL1; RGL2 and RGL3, by applying a modular design. The sensors are highly sensitive (in the low pm range), specific and dynamic. As a proof of concept, we have tested the applicability in three domains: the study of substrate specificity and activity of putative GA-oxidases, the characterization of GA transporters, and the use as a discrimination platform coupled to a GA agonists' chemical screening. This work demonstrates the development of a genetically encoded quantitative biosensor complementary to existing tools that allow the visualization of GA in planta."
Authors: Aneta Sienkiewicz, Małgorzata Krasowska, Małgorzata Kowczyk-Sadowy, Sławomir Obidziński, Alicja Piotrowska-Niczyporuk and Andrzej Bajguz
Scientific Reports (2024)
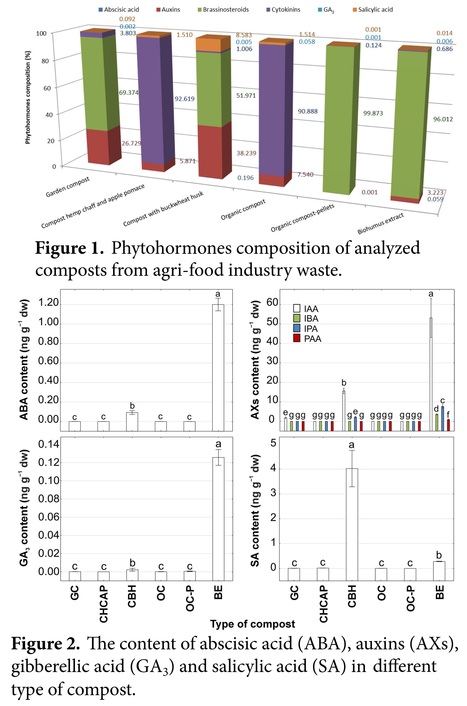
Abstract: "Utilizing the organic fraction of agri-food industry waste for fertilization represents one approach to waste management, with composting emerging as a popular method. Composts derived from this waste may contain plant hormones alongside primary macronutrients. This study aimed to evaluate the content of plant hormones in composts crafted from the organic fraction of agri-food industry waste. The presence of these substances was ascertained using liquid chromatography–mass spectrometry (LC-MS) analysis, applied to extracted samples from three composts produced in a bioreactor and three obtained from companies. The results indicate the presence of 35 compounds, which belong to six types of plant hormones: auxins, cytokinins, gibberellins, brassinosteroids, abscisic acid, and salicylic acid, in composts for the first time. The highest amount of plant hormones was noted in buckwheat husk and biohumus extract (35 compounds), and the lowest in hemp chaff and apple pomace (14 compounds). Brassinosteroids (e.g., brassinolide, 28-homobrassinolide, 24-epicastasterone, 24-epibrassinolide, and 28-norbrassinolide) and auxins (e.g., indolilo-3-acetic acid) are dominant. The highest concentration of total phytohormones was reported in biohumus extract (2026.42 ng g−1 dry weight), and the lowest in organic compost (0.18 ng g−1 dry weight)."
Authors: Guoling Guo, Lun Liu, Taijing Shen, Haozhe Wang, Shuqin Zhang, Yu Sun, Guoyu Xiong, Xiaomei Tang, Liwu Zhu and Bing Jia.
BMC Plant Biology (2024)
Abstract: "Background - Chlorophyll (Chl) is an agronomic trait associated with photosynthesis and yield. Gibberellin 2-oxidases (GA2oxs) have previously been shown to be involved in Chl accumulation. However, whether and how the PbrGA2ox proteins (PbrGA2oxs) mediate Chl accumulation in pear (Pyrus spp.) is scarce. Results - Here, we aimed to elucidate the role of the pear GA2ox gene family in Chl accumulation and the related underlying mechanisms. We isolated 13 PbrGA2ox genes (PbrGA2oxs) from the pear database and identified PbrGA2ox1 as a potential regulator of Chl accumulation. We found that transiently overexpressing PbrGA2ox1 in chlorotic pear leaves led to Chl accumulation, and PbrGA2ox1 silencing in normal pear leaves led to Chl degradation, as evident by the regreening and chlorosis phenomenon, respectively. Meanwhile, PbrGA2ox1-overexpressing (OE) tobacco plants discernably exhibited Chl built-up, as evidenced by significantly higher Pn and Fv/Fm. In addition, RNA sequencing (RNA-seq), physiological and biochemical investigations revealed an increase in abscisic acid (ABA), methyl jasmonate (MeJA), and salicylic acid (SA) concentrations and signaling pathways; a marked elevation in reducing and soluble sugar contents; and a marginal decline in the starch and sucrose levels in OE plants. Interestingly, PbrGA2ox1 overexpression did not prominently affect Chl synthesis. However, it indeed facilitated chloroplast development by increasing chloroplast number per cell and compacting the thylakoid granum stacks. These findings might jointly contribute to Chl accumulation in OE plants. Conclusion - Overall, our results suggested that GA2oxs accelerate Chl accumulation by stimulating chloroplast development and proved the potential of PbrGA2ox1 as a candidate gene for genetically breeding biofortified pear plants with a higher yield."
Authors: Yu-Lan Zhang, Li-Ying Wang, Yi Yang, Xu Zhao, Hong-Wei Zhu, Chen You, Ning Chen, Shuai-Jie Wei, Shu-Fen Li and Wu-Jun Gao.
The Plant Journal (2024)
Significance Statement: Using environmental factors such as hormones to induce sex conversion is an important approach to studying dioecy sex differentiation, which is a special character closely related to flowering, reproduction, and fruiting. In our research, exogenous GA3 can induce functional masculinization of individual spinach, and the GA-regulating pathway was established: GA-SpGAI-SpSTM-SpPI. This study provides new ideas and theoretical support for studying the sex differentiation mechanisms of spinach and other dioecious plants.
Abstract: "The sex of dioecious plants is mainly determined by genetic factors, but it can also be converted by environmental cues such as exogenous phytohormones. Gibberellic acids (GAs) are well-known inducers of flowering and sexual development, yet the pathway of gibberellin-induced sex conversion in dioecious spinach (Spinacia oleracea L.) remains elusive. Based on sex detection before and after GA3 application using T11A and SSR19 molecular markers, we confirmed and elevated the masculinization effect of GA on a single female plant through exogenous applications of GA3, showing complete conversion and functional stamens. Silencing of GIBBERELLIC ACID INSENSITIVE (SpGAI), a single DELLA family protein that is a central GA signaling repressor, results in similar masculinization. We also show that SpGAI can physically interact with the spinach KNOX transcription factor SHOOT MERISTEMLESS (SpSTM), which is a homolog of the flower meristem identity regulator STM in Arabidopsis. The silencing of SpSTM also masculinized female flowers in spinach. Furthermore, SpSTM could directly bind the intron of SpPI to repress SpPI expression in developing female flowers. Overall, our results suggest that GA induces a female masculinization process through the SpGAI-SpSTM-SpPI regulatory module in spinach. These insights may help to clarify the molecular mechanism underlying the sex conversion system in dioecious plants while also elucidating the physiological basis for the generation of unisexual flowers so as to establish dioecy in plants."
Authors: Xiaoxiao Li, Chuyu Lin, Chenghao Lan and Zeng Tao.
Journal of Experimental Botany (2024)
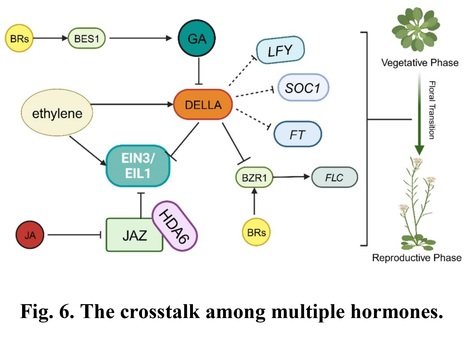
Abstract: "The timing of the developmental transition from the vegetative to the reproductive stages is critical for angiosperm and fine-tuned by the integration of endogenous factors and external environmental cues to ensure proper and successful reproduction. Plants have evolved sophisticated mechanisms to response to diverse environmental or stress signals, which may be mediated by plant hormones which coordinate their flowering time. Endogenous and exogenous phytohormones such as gibberellin (GA), auxin, cytokinin (CK), jasmonate (JA), abscisic acid (ABA), ethylene (ET), brassinosteroids (BR) and the cross-talk among them are critical for the precise regulating of flowering time. Recent studies on the model flowering plant Arabidopsis thaliana revealed that diverse transcription factors and epigenetic regulators play key roles in the phytohormones that regulate floral transition. This review aims to summarize current knowledge on the genetic and epigenetic mechanisms that underlying the phytohormone control of floral transition in Arabidopsis, offering insights into how these processes are regulated and their implications for plant biology."
|
Authors: Ping Huang, Jie Yang, Jiapeng Ke, Li Cai, Yingxiong Hu, Jun Ni, Chaoqiong Li, Zeng-Fu Xu and Mingyong Tang.
Plant Science (2024)
Highlights • GA4+7 and GA synthase gene JcGA20ox1 suppressed flowering in Jatropha, whereas the GA biosynthesis inhibitor paclobutrazol and GA degradation gene JcGA2ox6 promoted flowering. • JcGA20ox1 decreased the expression of JcFT; while, the inhibition of floral transition by JcGA20ox1 could restored by JcFT, and the seed yield suppression phenotype could partially restored by JcFT. • JcGA20ox1 also decreased JiFT expression and delayed floral transition in J. curcas × J. integerrima hybrids.
Abstract: "Jatropha curcas is a perennial oil-seed plant with vigorous vegetative growth but relatively poor reproductive growth and low seed yield. Gibberellins (GA) promote flowering in most annual plants, but inhibit flower initiation in many woody plants, including J. curcas. However, the underlying mechanisms of GA inhibition on flowering in perennial woody plants remain unclear. Here, we found that overexpression of the gibberellin biosynthesis gene JcGA20ox1 delays flowering in J. curcas and in the J. curcas × J. integerrima hybrids. Consistent with this finding, overexpression of the GA catabolic gene JcGA2ox6 induces early flowering in J. curcas. qRTsingle bondPCR revealed that the inhibition of floral transition by overexpressing JcGA20ox1 resulted from the decreased of JcFT and other flowering related genes, which was restored by overexpressing JcFT in J. curcas. Overexpression of JcGA20ox1 or JcGA2ox6 reduced seed yield, but overexpression of JcFT significantly increased seed yield. Furthermore, hybridization experiments showed that the reduction in seed yield caused by overexpression of JcGA20ox1 or JcGA2ox6 was partially restored by the overexpression of JcFT. In addition, JcGA20ox1, JcGA2ox6 and JcFT were also found to be involved in the regulation of seed oil content and endosperm development. In conclusion, our study revealed the inhibitory effect of GA on flowering in J. curcas is mediated through JcFT and demonstrated the effects of JcGA20ox1, JcGA2ox6 and JcFT on agronomic traits. This study also indicates the potential value of GA metabolism genes and JcFT in breeding new varieties of woody oil-seed plants."
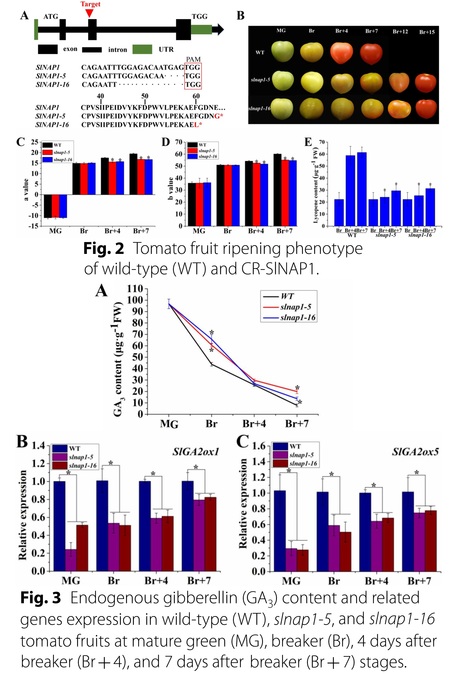
Authors: Changxia Li, Xuemei Hou, Zongxi Zhao, Huwei Liu, Panpan Huang, Meimei Shi, Xuetong Wu, Rong Gao, Zhiya Liu, Lijuan Wei, Yihua Li and Weibiao Liao.
Cellular & Molecular Biology Letters (2024)
Abstract: "In tomato (Solanum lycopersicum), the ripening of fruit is regulated by the selective expression of ripening-related genes, and this procedure is controlled by transcription factors (TFs). In the various plant-specific TF families, the no apical meristem (NAM), Arabidopsis thaliana activating factor 1/2 (ATAF1/2), and cup-shaped cotyledon 2 (CUC2; NAC) TF family stands out and plays a significant function in plant physiological activities, such as fruit ripening (FR). Despite the numerous genes of NAC found in the tomato genome, limited information is available on the effects of NAC members on FR, and there is also a lack of studies on their target genes. In this research, we focus on SlNAP1, which is a NAC TF that positively influences the FR of tomato. By employing CRISPR/Cas9 technology, compared with the wild type (WT), we generated slnap1 mutants and observed a delay in the ethylene production and color change of fruits. We employed the yeast one-hybrid (Y1H) and dual-luciferase reporter (DLR) assays to confirm that SlNAP1 directly binds to the promoters of two crucial genes involved in gibberellin (GA) degradation, namely SlGA2ox1 and SlGA2ox5, thus activating their expression. Furthermore, through a yeast two-hybrid (Y2H), bimolecular fluorescence complementation (BIFC) and luciferase (LUC) assays, we established an interaction between SlNAP1 and SlGID1. Hence, our findings suggest that SlNAP1 regulates FR positively by activating the GA degradation genes directly. Additionally, the interaction between SlNAP1 and SlGID1 may play a role in SlNAP1-induced FR. Overall, our study provides important insights into the molecular mechanisms through which NAC TFs regulate tomato FR via the GA pathway."
Authors: Wanmin Wang, Zizhao Xie, Yuanyuan Wu, Ying Sun, Chenghang Zhan, Liang Jin and Junli Huang.
Environmental and Experimental Botany (2024)
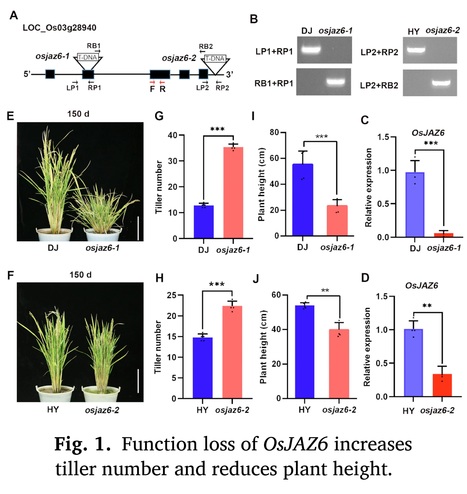
Highlights: • OsJAZ6 modulates rice tillering and drought response by integrating JA with GA signaling. • OsJAZ6 controls the tiller bud growth but not formation. • OsJAZ6 interacts with SLR1 to promote its degradation, which further destabilizes MOC1. • OsJAZ6 and SLR1 have opposite functions in regulating rice tiller bud growth and drought tolerance.
Abstract: "Jasmonic acid (JA) plays crucial functions during plant growth and stress response, but its roles and regulatory mechanism in plant branching remain largely unknown. Rice basal branching (tillering) is an essential agronomic trait that affects crop production. Here, we report that OsJAZ6, the repressor of JA signaling, negatively modulates rice tillering and drought stress tolerance. Loss-of-function mutants of OsJAZ6 exhibit a significant increase in tiller number, while OsJAZ6ΔJas-overexpression lines produce fewer tillers than wild-type plants. Further investigations show that function loss of OsJAZ6 promotes the tiller bud growth rather than formation. Mechanistic studies show that OsJAZ6 interacts with rice DELLA/SLR1 (SLENDER RICE 1), a transcription repressor of gibberellin (GA) signaling, and the interaction promotes SLR1 degradation, which further facilitates the degradation of rice tillering regulator MOC1 (MONOCULM 1), thereby inhibiting the tiller bud growth. In agreement, the slr1 mutant exhibits fewer tillers than wild type. Consistently, application of JA promotes the growth of tiller bud and thus increases the tiller number, while GA treatment results in opposite result. Meanwhile, osjaz6 mutants display enhanced drought tolerance, coupled with increased JA sensitivity, while the slr1 mutant shows the reverse behavior. Collectively, our data demonstrate that OsJAZ6 negatively modulates rice tillering as well as drought stress tolerance by destabilizing SLR1 protein. Our data shed light on the regulatory mechanism of controlling the tiller development and drought stress response in rice by the JA-OsJAZ6-SLR1 module."
Authors: Pauline Raimondeau, Sayam Ksouda, William Marande, Anne-Laure Fuchs, Hervé Gryta, Anthony Theron, Aurore Puyoou, Julia Dupin, Pierre-Olivier Cheptou, Sonia Vautrin, Sophie Valière, Sophie Manzi, Djamel Baali-Cherif, Jérôme Chave, Pascal-Antoine Christin and Guillaume Besnard.
Current Biology (2024)
Editor's view: Raimondeau et al. unveil the genetic determinants of the unique self-incompatibility system of olives. This initial discovery leads them to identify the genetic controls of distyly within the family and to uncover a genetic overlap between the two systems, rooted in shared ancestry.
Highlights: • Homomorphic self-incompatibility in olives is associated with a hemizygous region • A homologous hemizygous region is also involved in controlling distyly in jasmine • The duplication of this DNA segment is ancestral in the family • Hormonal regulation is central to the production of self-incompatibility phenotypes
Abstract: "Self-incompatibility (SI) has evolved independently multiple times and prevents self-fertilization in hermaphrodite angiosperms. Several groups of Oleaceae such as jasmines exhibit distylous flowers, with two compatibility groups each associated with a specific floral morph.1 Other Oleaceae species in the olive tribe have two compatibility groups without associated morphological variation.2,3,4,5 The genetic basis of both homomorphic and dimorphic SI systems in Oleaceae is unknown. By comparing genomic sequences of three olive subspecies (Olea europaea) belonging to the two compatibility groups, we first locate the genetic determinants of SI within a 700-kb hemizygous region present only in one compatibility group. We then demonstrate that the homologous hemizygous region also controls distyly in jasmine. Phylogenetic analyses support a common origin of both systems, following a segmental genomic duplication in a common ancestor. Examination of the gene content of the hemizygous region in different jasmine and olive species suggests that the mechanisms determining compatibility groups and floral phenotypes (whether homomorphic or dimorphic) in Oleaceae rely on the presence/absence of two genes involved in gibberellin and brassinosteroid regulation."
Authors: Jiujun Du, Hantian Wei, Xueqin Song, Lei Zhang and Jianjun Hu.
Plant Science (2024)
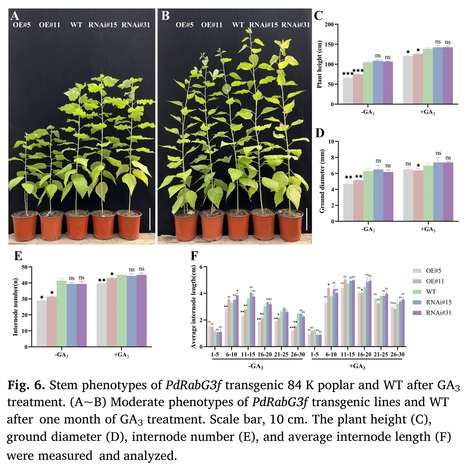
Highlights: • PdRabG3f inhibited the vertical elongation of poplar by suppressing internode elongation. • PdRabG3f inhibited xylem development by suppressing cambium division and differentiation. • PdRabG3f interfered with gibberellin-mediated internode elongation and xylem developing in poplar.
Abstract: "As a member of the small GTPases family, Rab GTPases play a key role in specifying transport pathways in the intracellular membrane trafficking system and are involved in plant growth and development. By quantitative trait locus (QTL) mapping, PdRabG3f was identified as a candidate gene associated with shoot height in a hybrid offspring of Populus deltoides ‘Danhong’ × Populus simonii ‘Tongliao1’. PdRabG3f localized to the nucleus, endoplasmic reticulum and tonoplast and was primarily expressed in the xylem and cambium. Overexpression of PdRabG3f in Populus alba × Populus glandulosa (84 K poplar) had inhibitory effects on vertical and radical growth. In the transgenic lines, there were evident changes in the levels of 15 gibberellin (GA) derivatives, and the application of exogenous GA3 partially restored the phenotypes mediated by GAs deficiency. The interaction between PdRabG3f and RIC4, which was the GA-responsive factor, provided additional explanation for PdRabG3f's inhibitory effect on poplar growth. RNA-seq analysis revealed differentially expressed genes (DEGs) associated with cell wall, xylem, and gibberellin response. PdRabG3f interfering endogenous GAs levels in poplar might involve the participation of MYBs and ultimately affected internode elongation and xylem development. This study provides a potential mechanism for gibberellin-mediated regulation of plant growth through Rab GTPases."
Authors: Nguyen Hoai Nguyen, Phuong Thi Bich Ho and Linh Thi Truc Le.
Plant Reproduction (2024)
Key message: This review provides a thorough and comprehensive perspective on the topic of cucumber sexual expression. Specifically, insights into sex expression mediated by pathways other than ethylene are highlighted.
Abstract: "Cucumber (Cucumis sativus L.) is a common and important commercial crop that is cultivated and consumed worldwide. Additionally, this species is commonly used as a model for investigating plant sex expression. Cucumbers exhibit a variety of floral arrangements, comprising male, female, and hermaphroditic (bisexual) flowers. Generally, cucumber plants that produce female flowers are typically preferred due to their significant impact on the overall output. Various environmental conditions, such as temperature, light quality, and photoperiod, have been also shown to influence the sex expression in this species. Multiple lines of evidence indicate that ethylene and its biosynthesis genes are crucial in regulating cucumber sex expression. Gibberellins, another well-known phytohormone, can similarly influence cucumber sex expression via an ethylene-independent route. Further studies employing the next-generation sequencing technology also visualized a deeper slice of the molecular mechanism such as the role of the cell cycle program in the cucumber sex expression. This review aims to provide an overview of the sex expression of cucumber including its underlying molecular mechanism and regulatory aspects based on recent investigations."
Authors: Junpeng Niu, Mingzhen Xu, Na Zong, Jia Sun, Lei Zhao and Wei Hui.
Physiologia Plantarum (2024)
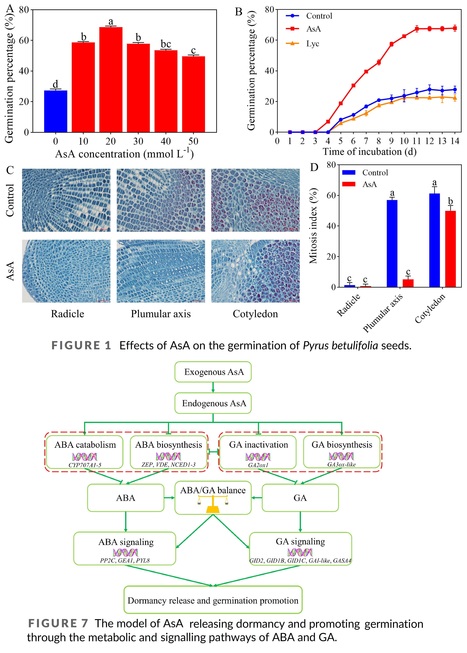
Abstract: "Seed dormancy is an important life history state in which intact viable seeds delay or prevent germination under suitable conditions. Ascorbic acid (AsA) acts as a small molecule antioxidant, and breaking seed dormancy and promoting subsequent growth are among its numerous functions. In this study, a germination test using Pyrus betulifolia seeds treated with exogenous AsA or AsA synthesis inhibitor lycorine (Lyc) and water absorption was conducted. The results indicated that AsA released dormancy and increased germination and 20 mmol L−1 AsA promoted cell division, whereas Lyc reduced germination. Seed germination showed typical three phases of water absorption; and seeds at five key time points were sampled for transcriptome analysis. It revealed that multiple pathways were involved in breaking dormancy and promoting germination through transcriptome data, and 12 differentially expressed genes (DEGs) related to the metabolism and signal transduction of abscisic acid (ABA) and gibberellins (GA) were verified by subsequent RT-qPCR. For metabolites, exogenous AsA increased endogenous AsA and GA3 but reduced ABA and the ABA/GA3 ratio. In addition, three genes regulating ABA synthesis were downregulated by AsA, while five genes mediating ABA degradation were upregulated. Taken together, AsA regulates the pathways associated with ABA and GA synthesis, catalysis, and signal transduction, with subsequent reduction in ABA and increase in GA and further the balance of ABA/GA, ultimately releasing dormancy and promoting germination."
Authors: Clare Hurst and Miriam L. Gifford.
Development (2024)
Excepts: "Legumes form a symbiotic relationship with bacteria called rhizobia. The rhizobia fix atmospheric dinitrogen, which is used by the plant as a nitrogen resource, and take up molecules, including carbon compounds, from the plant. Rhizobia are housed in nodules that develop on roots after a series of orchestrated stages that are triggered by the perception and entry of rhizobia at root hairs (reviewed by Luo et al., 2023). Study of this relationship and the mechanisms that govern nodule formation and inhibition may reveal targets to promote enhanced nodulation efficiency and increased nitrogen acquisition for the plant. Phytohormones have been shown to play a key role in both nodule formation and inhibition; therefore, a better understanding of hormonal regulatory activity could provide options for such enhancement. To address this, Drapek and colleagues (2023 preprint) focus on the dynamics of gibberellin (GA), a phytohormone previously identified as both a positive and negative regulator of nodulation (Rizza et al., 2017; Fonouni-Farde et al., 2016).
"In conclusion, this study analysed the spatial and temporal accumulation of GA, the interactions between GA and key nodule identity regulatory genes, and tested the importance of GA biosynthesis enzymes in nodule formation. The work provides strong evidence that GA is a potential determinator between lateral root formation and nodulation, and helps to explain previous contradictory findings related to the importance of GA, as well as cytokinin, in these processes."
Authors: Huibin Xu, Fuxiang Wang, Rebecca Njeri Damari, Xiaofeng Chen and Zhongyuan Lin.
Molecular Breeding (2024)
Abstract: "Seed germination is a vital step in the life cycle of a plant, playing a significant role in seedling establishment and crop yield potential. It is also an important factor in the conservation of plant germplasm resources. This complex process is influenced by a myriad of factors, including environmental conditions, the genetic makeup of the seed, and endogenous hormones. The perception of these environmental signals triggers a cascade of intricate signal transduction events that determine whether a seed germinates or remains dormant. Despite considerable progress in uncovering the molecular mechanisms governing these processes, many questions remain unanswered. In this review, we summarize the current progress in the molecular mechanisms underlying the perception of environmental signals and consequent signal transduction during seed germination, and discuss questions that need to be addressed to better understand the process of seed germination and develop novel strategies for germplasm improvement."
Authors: Barbora Ndreca, Alison Huttly, Sajida Bibi, Carlos Bayon, George Lund, Joshua Ham, Rocío Alarcón-Reverte, John Addy, Danuše Tarkowská, Stephen Pearce, Peter Hedden, Stephen G. Thomas and Andrew L. Phillips.
Research Square (2024)
Abstract: "Background - Semi-dwarfing alleles are used widely in cereals to confer improved lodging resistance and assimilate partitioning. The most widely deployed semi-dwarfing alleles in rice and barley encode the gibberellin (GA)-biosynthetic enzyme GA 20-OXIDASE2 (GA20OX2). The hexaploid wheat genome carries three homoeologous copies of GA20OX2, and because of functional redundancy, loss-of-function alleles of a single homoeologue would not be selected in wheat breeding programmes. Instead, approximately 70% of wheat cultivars carry gain-of-function mutations in REDUCED HEIGHT 1 (RHT1) genes that encode negative growth regulators and are degraded in response to GA. Semi-dwarf Rht-B1b or Rht-D1b alleles encode proteins that are insensitive to GA-mediated degradation. However, because RHT1 is expressed ubiquitously these alleles have pleiotropic effects that confer undesirable traits in some environments. Results - We have applied reverse genetics to combine loss-of-function alleles in all three homoeologues of wheat GA20OX2 and its paralogue GA20OX1 and evaluated their performance in three years of field trials. ga20ox1 mutants exhibited a mild height reduction (approximately 3%) suggesting GA20OX1 plays a minor role in stem elongation in wheat. ga20ox2 mutants have reduced GA1 content and are 12-32% shorter than their wild-type segregants, comparable to the effect of the Rht-D1b ‘Green Revolution’ allele. The ga20ox2 mutants showed no significant negative effects on yield components, although these alleles should be evaluated in different genetic backgrounds and environments. Conclusions - Our study demonstrates that induced mutagenesis can expand genetic variation in polyploid crops to uncover novel alleles and that mutations in GA20OX2 could have utility in wheat breeding as alternative semi-dwarfing alleles.
Authors: Peng Zeng, Ting Xie, Jiaxin Shen, Taokai Liang, Lu Yin, Kexin Liu, Ying He, Mingming Chen, Haijuan Tang, Sunlu Chen, Sergey Shabala, Hongsheng Zhang and Jinping Cheng.
Journal of Integrative Plant Biology (2024)
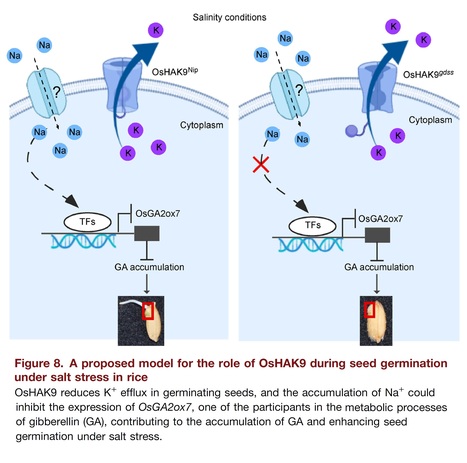
Abstract: "Soil salinity has a major impact on rice seed germination, severely limiting rice production. Herein, a rice germination defective mutant under salt stress (gdss) was identified by using chemical mutagenesis. The GDSS gene was detected via MutMap and shown to encode potassium transporter OsHAK9. Phenotypic analysis of complementation and mutant lines demonstrated that OsHAK9 was an essential regulator responsible for seed germination under salt stress. OsHAK9 is highly expressed in germinating seed embryos. Ion contents and non-invasive micro-test technology results showed that OsHAK9 restricted K+ efflux in salt-exposed germinating seeds for the balance of K+/Na+. Disruption of OsHAK9 significantly reduced gibberellin 4 (GA4) levels, and the germination defective phenotype of oshak9a was partly rescued by exogenous GA3 treatment under salt stress. RNA sequencing (RNA-seq) and real-time quantitative polymerase chain reaction analysis demonstrated that the disruption of OsHAK9 improved the GA-deactivated gene OsGA2ox7 expression in germinating seeds under salt stress, and the expression of OsGA2ox7 was significantly inhibited by salt stress. Null mutants of OsGA2ox7 created using clustered, regularly interspaced, short palindromic repeat (CRISPR)/CRISPR-associated nuclease 9 approach displayed a dramatically increased seed germination ability under salt stress. Overall, our results highlight that OsHAK9 regulates seed germination performance under salt stress involving preventing GA degradation by mediating OsGA2ox7, which provides a novel clue about the relationship between GA and OsHAKs in rice."
Authors: Enyang Mei, Mingliang He, Min Xu, Jiaqi Tang, Jiali Liu, Yingxiang Liu, Zhipeng Hong, Xiufeng Li, Zhenyu Wang, Qingjie Guan, Xiaojie Tian and Qingyun Bu.
Journal of Integrative Plant Biology (2024)
Abstract: "Panicle exsertion is one of the crucial agronomic traits in rice (Oryza sativa). Shortening of panicle exsertion often leads to panicle enclosure and severely reduces seed production. Gibberellin (GA) plays important roles in regulating panicle exsertion. However, the underlying mechanism and the relative regulatory network remain elusive. Here, we characterized the oswrky78 mutant showing severe panicle enclosure, and found that the defect of oswrky78 is caused by decreased bioactive GA contents. Biochemical analysis demonstrates that OsWRKY78 can directly activate GA biosynthesis and indirectly suppress GA metabolism. Moreover, we found OsWRKY78 can interact with and be phosphorylated by mitogen-activated protein kinase (MAPK) kinase OsMAPK6, and this phosphorylation can enhance OsWRKY78 stability and is necessary for its biological function. Taken together, these results not only reveal the critical function of OsWRKY78, but also reveal its mechanism via mediating crosstalk between MAPK and the GA signaling pathway in regulating panicle exsertion."
|



 Your new post is loading...
Your new post is loading...